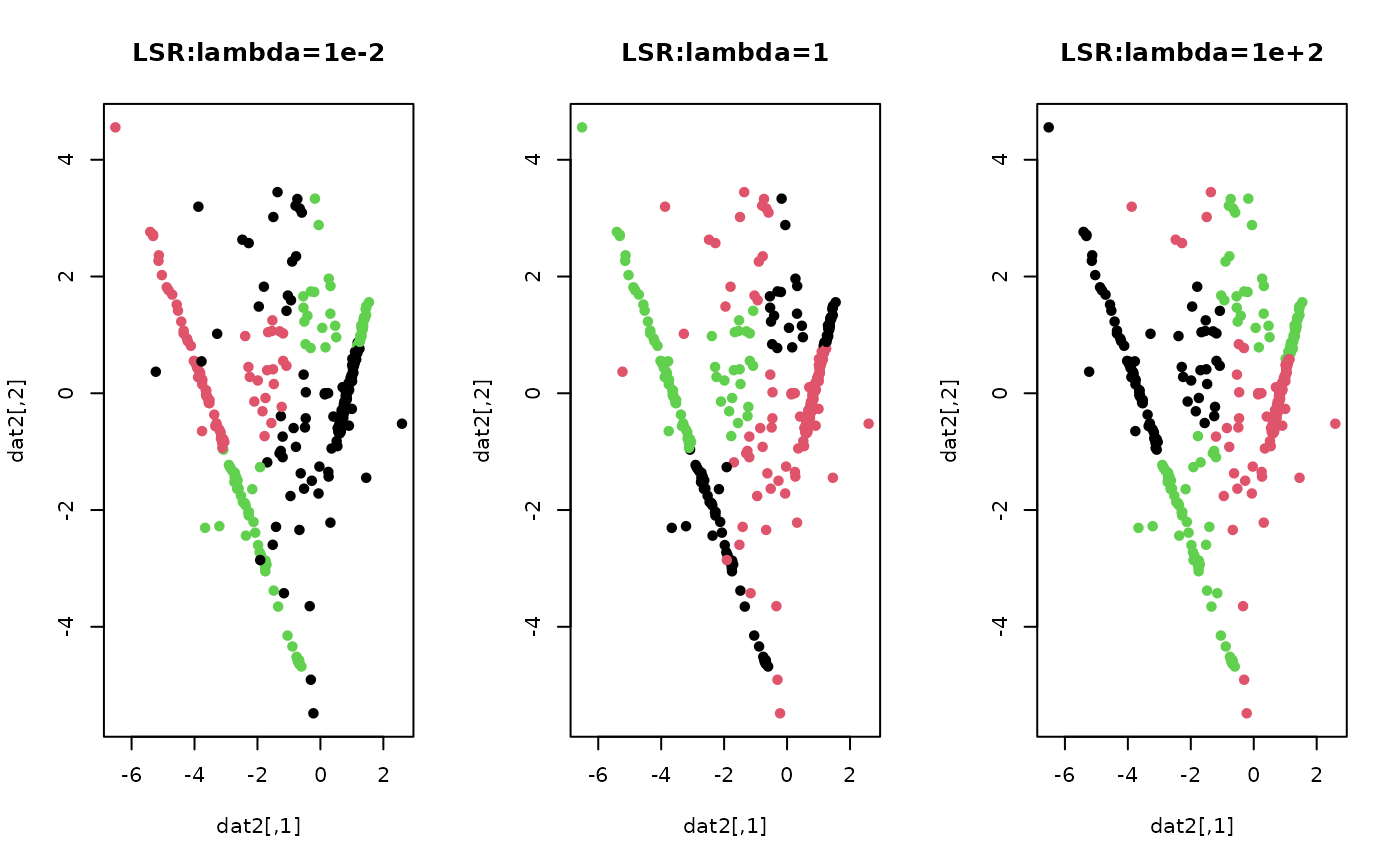For the subspace clustering, traditional method of least squares regression
is used to build coefficient matrix that reconstructs the data point by solving
$$\textrm{min}_Z \|X-XZ\|_F^2 + \lambda \|Z\|_F \textrm{ such that }diag(Z)=0$$
where \(X\in\mathbf{R}^{p\times n}\) is a column-stacked data matrix.
As seen from the equation, we use a denoising version controlled by \(\lambda\) and
provide an option to abide by the constraint \(diag(Z)=0\) by zerodiag parameter.
LSR(data, k = 2, lambda = 1e-05, zerodiag = TRUE)
Arguments
| data | an \((n\times p)\) matrix of row-stacked observations. |
|---|---|
| k | the number of clusters (default: 2). |
| lambda | regularization parameter (default: 1e-5). |
| zerodiag | a logical; |
Value
a named list of S3 class T4cluster containing
- cluster
a length-\(n\) vector of class labels (from \(1:k\)).
- algorithm
name of the algorithm.
References
Lu C, Min H, Zhao Z, Zhu L, Huang D, Yan S (2012). “Robust and Efficient Subspace Segmentation via Least Squares Regression.” In Hutchison D, Kanade T, Kittler J, Kleinberg JM, Mattern F, Mitchell JC, Naor M, Nierstrasz O, Pandu Rangan C, Steffen B, Sudan M, Terzopoulos D, Tygar D, Vardi MY, Weikum G, Fitzgibbon A, Lazebnik S, Perona P, Sato Y, Schmid C (eds.), Computer Vision -ECCV 2012, volume 7578, 347--360. Springer Berlin Heidelberg, Berlin, Heidelberg. ISBN 978-3-642-33785-7 978-3-642-33786-4.
Examples
# \donttest{ ## generate a toy example set.seed(10) tester = genLP(n=100, nl=2, np=1, iso.var=0.1) data = tester$data label = tester$class ## do PCA for data reduction proj = base::eigen(stats::cov(data))$vectors[,1:2] dat2 = data%*%proj ## run LSR for k=3 with different lambda values out1 = LSR(data, k=3, lambda=1e-2) out2 = LSR(data, k=3, lambda=1) out3 = LSR(data, k=3, lambda=1e+2) ## extract label information lab1 = out1$cluster lab2 = out2$cluster lab3 = out3$cluster ## visualize opar <- par(no.readonly=TRUE) par(mfrow=c(1,3)) plot(dat2, pch=19, cex=0.9, col=lab1, main="LSR:lambda=1e-2") plot(dat2, pch=19, cex=0.9, col=lab2, main="LSR:lambda=1") plot(dat2, pch=19, cex=0.9, col=lab3, main="LSR:lambda=1e+2")