Orthogonal Neighborhood Preserving Projection (ONPP) is an unsupervised linear dimension reduction method. It constructs a weighted data graph from LLE method. Also, it develops LPP method by preserving the structure of local neighborhoods.
Arguments
- X
an \((n\times p)\) matrix or data frame whose rows are observations and columns represent independent variables.
- ndim
an integer-valued target dimension.
- type
a vector of neighborhood graph construction. Following types are supported;
c("knn",k),c("enn",radius), andc("proportion",ratio). Default isc("proportion",0.1), connecting about 1/10 of nearest data points among all data points. See alsoaux.graphnbdfor more details.- preprocess
an additional option for preprocessing the data. Default is "center". See also
aux.preprocessfor more details.
Value
a named list containing
- Y
an \((n\times ndim)\) matrix whose rows are embedded observations.
- trfinfo
a list containing information for out-of-sample prediction.
- projection
a \((p\times ndim)\) whose columns are basis for projection.
References
Kokiopoulou E, Saad Y (2007). “Orthogonal Neighborhood Preserving Projections: A Projection-Based Dimensionality Reduction Technique.” IEEE Transactions on Pattern Analysis and Machine Intelligence, 29(12), 2143–2156.
Examples
## use iris data
data(iris)
set.seed(100)
subid = sample(1:150, 50)
X = as.matrix(iris[subid,1:4])
label = as.factor(iris[subid,5])
## try different numbers for neighborhood size
out1 = do.onpp(X, type=c("proportion",0.10))
out2 = do.onpp(X, type=c("proportion",0.25))
out3 = do.onpp(X, type=c("proportion",0.50))
## visualize
opar <- par(no.readonly=TRUE)
par(mfrow=c(1,3))
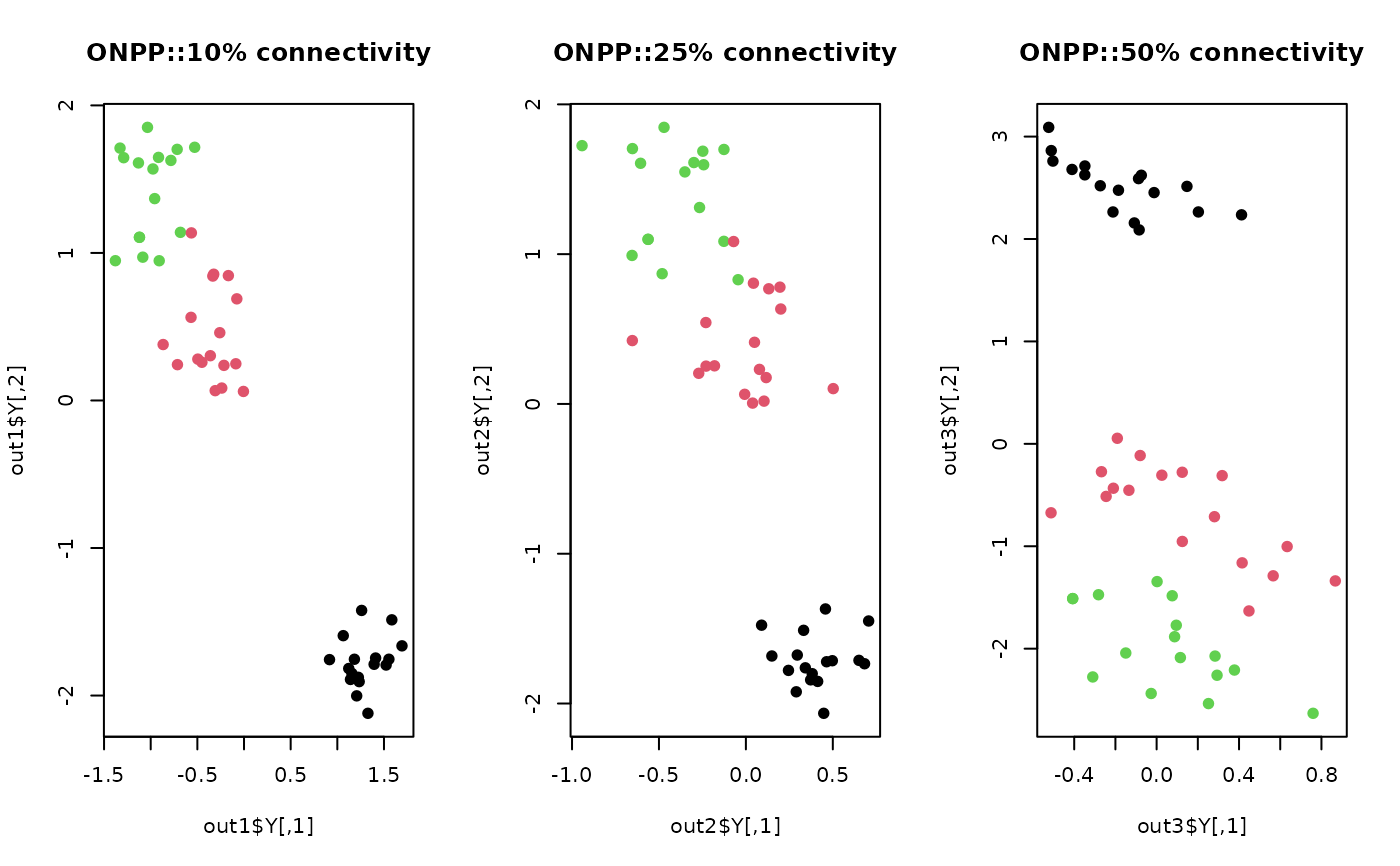
plot(out1$Y, pch=19, col=label, main="ONPP::10% connectivity")
plot(out2$Y, pch=19, col=label, main="ONPP::25% connectivity")
plot(out3$Y, pch=19, col=label, main="ONPP::50% connectivity")
 par(opar)
par(opar)