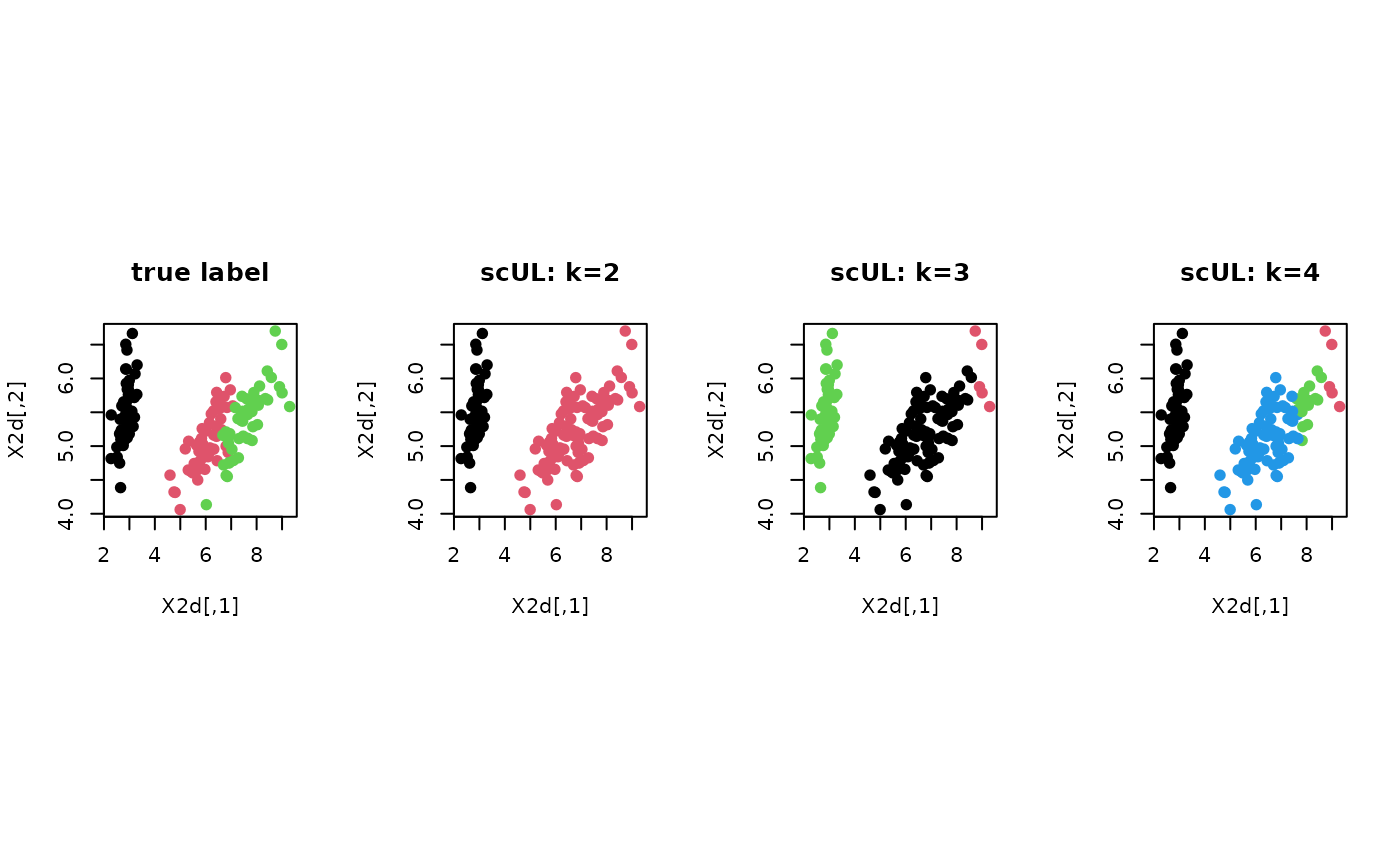The version of Shi and Malik first constructs the affinity matrix $$A_{ij} = \exp(-d(x_i, d_j)^2 / \sigma^2)$$ where \(\sigma\) is a common bandwidth parameter and performs k-means (or possibly, GMM) clustering on the row-space of eigenvectors for the unnormalized graph laplacian matrix $$L=D-A$$.
scUL(data, k = 2, sigma = 1, ...)
Arguments
| data | an \((n\times p)\) matrix of row-stacked observations or S3 |
|---|---|
| k | the number of clusters (default: 2). |
| sigma | bandwidth parameter (default: 1). |
| ... | extra parameters including
|
Value
a named list of S3 class T4cluster containing
- cluster
a length-\(n\) vector of class labels (from \(1:k\)).
- eigval
eigenvalues of the graph laplacian's spectral decomposition.
- embeds
an \((n\times k)\) low-dimensional embedding.
- algorithm
name of the algorithm.
References
von Luxburg U (2007). “A Tutorial on Spectral Clustering.” Statistics and Computing, 17(4), 395--416. ISSN 0960-3174, 1573-1375.
Examples
# ------------------------------------------------------------- # clustering with 'iris' dataset # ------------------------------------------------------------- ## PREPARE data(iris) X = as.matrix(iris[,1:4]) lab = as.integer(as.factor(iris[,5])) ## EMBEDDING WITH PCA X2d = Rdimtools::do.pca(X, ndim=2)$Y ## CLUSTERING WITH DIFFERENT K VALUES cl2 = scUL(X, k=2)$cluster cl3 = scUL(X, k=3)$cluster cl4 = scUL(X, k=4)$cluster ## VISUALIZATION opar <- par(no.readonly=TRUE) par(mfrow=c(1,4), pty="s") plot(X2d, col=lab, pch=19, main="true label") plot(X2d, col=cl2, pch=19, main="scUL: k=2") plot(X2d, col=cl3, pch=19, main="scUL: k=3") plot(X2d, col=cl4, pch=19, main="scUL: k=4")