Metric MDS is a nonlinear method that is solved iteratively. We adopt a well-known SMACOF algorithm for updates with uniform weights over all pairwise distances after initializing the low-dimensional configuration via classical MDS.
do.mmds(X, ndim = 2, ...)Arguments
- X
an \((n\times p)\) matrix or data frame whose rows are observations and columns represent independent variables.
- ndim
an integer-valued target dimension (default: 2).
- ...
extra parameters including
- maxiter
maximum number of iterations for metric MDS updates (default: 100).
- abstol
stopping criterion for metric MDS iterations (default: 1e-8).
Value
a named Rdimtools S3 object containing
- Y
an \((n\times ndim)\) matrix whose rows are embedded observations.
- algorithm
name of the algorithm.
References
Leeuw JD, Barra IJR, Brodeau F, Romier G, (eds BVC (1977). “Applications of Convex Analysis to Multidimensional Scaling.” In Recent Developments in Statistics, 133--146.
Borg I, Groenen PJF (2010). Modern Multidimensional Scaling: Theory and Applications. Springer New York, New York, NY. ISBN 978-1-4419-2046-1 978-0-387-28981-6.
Examples
# \donttest{
## load iris data
data(iris)
X = as.matrix(iris[,1:4])
lab = as.factor(iris[,5])
## compare with other methods
pca2d <- do.pca(X, ndim=2)
cmd2d <- do.mds(X, ndim=2)
mmd2d <- do.mmds(X, ndim=2)
## Visualize
opar <- par(no.readonly=TRUE)
par(mfrow=c(1,3))
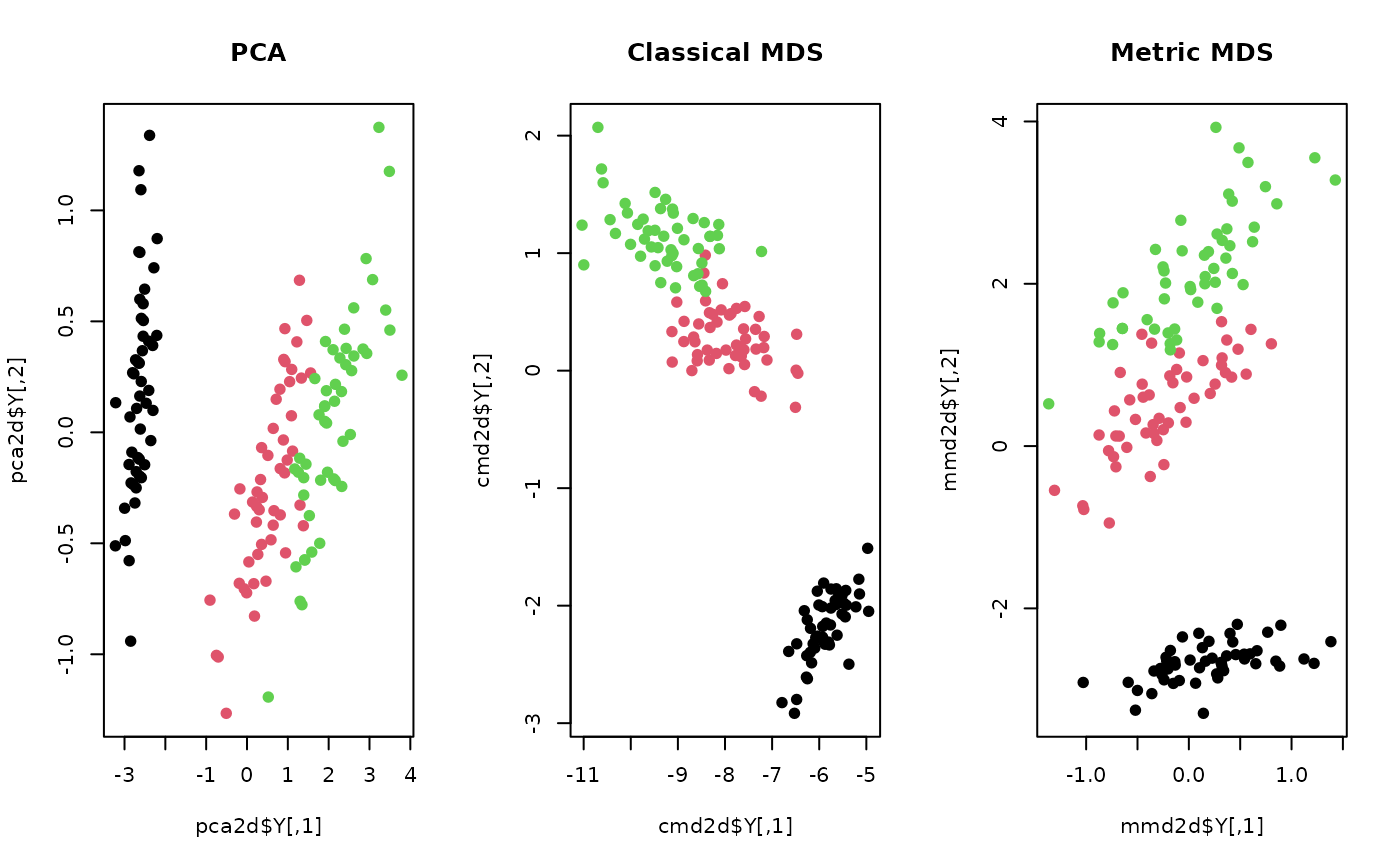
plot(pca2d$Y, col=lab, pch=19, main="PCA")
plot(cmd2d$Y, col=lab, pch=19, main="Classical MDS")
plot(mmd2d$Y, col=lab, pch=19, main="Metric MDS")
 par(opar)
# }
par(opar)
# }