do.dm discovers low-dimensional manifold structure embedded in high-dimensional
data space using Diffusion Maps (DM). It exploits diffusion process and distances in data space to find
equivalent representations in low-dimensional space.
do.dm(
X,
ndim = 2,
preprocess = c("null", "center", "scale", "cscale", "decorrelate", "whiten"),
bandwidth = 1,
timescale = 1,
multiscale = FALSE
)Arguments
- X
an \((n\times p)\) matrix or data frame whose rows are observations and columns represent independent variables.
- ndim
an integer-valued target dimension.
- preprocess
an additional option for preprocessing the data. Default is "null". See also
aux.preprocessfor more details.- bandwidth
a scaling parameter for diffusion kernel. Default is 1 and should be a nonnegative real number.
- timescale
a target scale whose value represents behavior of heat kernels at time t. Default is 1 and should be a positive real number.
- multiscale
logical;
FALSEis to use the fixedtimescalevalue,TRUEto ignore the given value.
Value
a named list containing
- Y
an \((n\times ndim)\) matrix whose rows are embedded observations.
- trfinfo
a list containing information for out-of-sample prediction.
- eigvals
a vector of eigenvalues for Markov transition matrix.
References
Nadler B, Lafon S, Coifman RR, Kevrekidis IG (2005). “Diffusion Maps, Spectral Clustering and Eigenfunctions of Fokker-Planck Operators.” In Proceedings of the 18th International Conference on Neural Information Processing Systems, NIPS'05, 955–962.
Coifman RR, Lafon S (2006). “Diffusion Maps.” Applied and Computational Harmonic Analysis, 21(1), 5–30.
Examples
# \donttest{
## load iris data
data(iris)
set.seed(100)
subid = sample(1:150,50)
X = as.matrix(iris[subid,1:4])
label = as.factor(iris[subid,5])
## compare different bandwidths
out1 <- do.dm(X,bandwidth=10)
out2 <- do.dm(X,bandwidth=100)
out3 <- do.dm(X,bandwidth=1000)
## visualize
opar <- par(no.readonly=TRUE)
par(mfrow=c(1,3))
plot(out1$Y, pch=19, col=label, main="DM::bandwidth=10")
plot(out2$Y, pch=19, col=label, main="DM::bandwidth=100")
plot(out3$Y, pch=19, col=label, main="DM::bandwidth=1000")
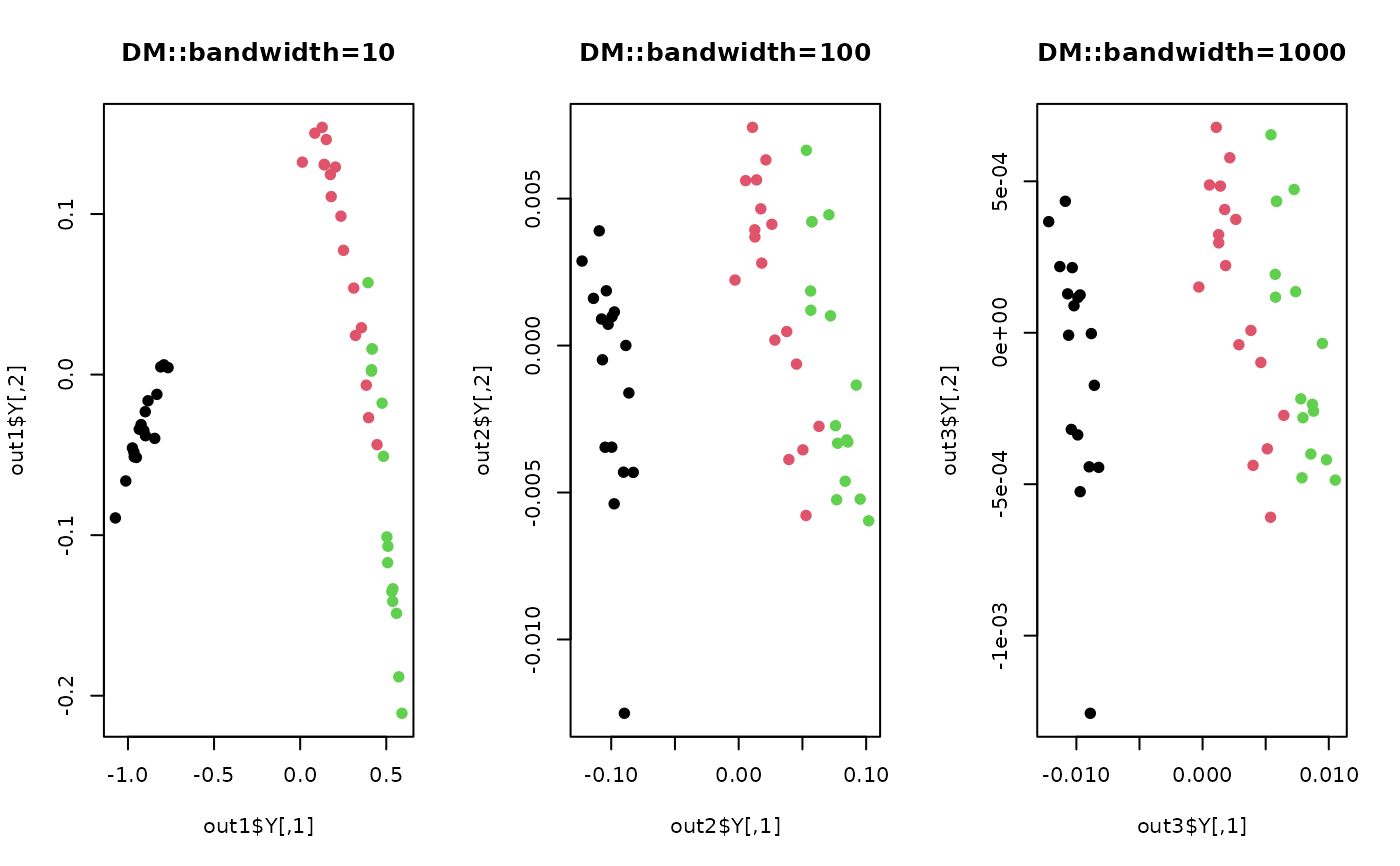 par(opar)
# }
par(opar)
# }