This unsupervised feature selection method is based on self-expression model, which means that the cost function involves difference in self-representation. It does not explicitly require learning the clusterings and different features are weighted individually based on their relative importance. The cost function involves two penalties, sparsity and preservation of local structure.
do.spufs(
X,
ndim = 2,
preprocess = c("null", "center", "scale", "cscale", "whiten", "decorrelate"),
alpha = 1,
beta = 1,
bandwidth = 1
)Arguments
- X
an \((n\times p)\) matrix or data frame whose rows are observations and columns represent independent variables.
- ndim
an integer-valued target dimension.
- preprocess
an additional option for preprocessing the data. Default is "null". See also
aux.preprocessfor more details.- alpha
nonnegative number to control sparsity in rows of matrix of representation coefficients.
- beta
nonnegative number to control the degree of local-structure preservation.
- bandwidth
positive number for Gaussian kernel bandwidth to define similarity.
Value
a named list containing
- Y
an \((n\times ndim)\) matrix whose rows are embedded observations.
- featidx
a length-\(ndim\) vector of indices with highest scores.
- trfinfo
a list containing information for out-of-sample prediction.
- projection
a \((p\times ndim)\) whose columns are basis for projection.
References
Lu Q, Li X, Dong Y (2018). “Structure Preserving Unsupervised Feature Selection.” Neurocomputing, 301, 36--45.
Examples
## use iris data
data(iris)
set.seed(100)
subid = sample(1:150, 50)
X = as.matrix(iris[subid,1:4])
label = as.factor(iris[subid,5])
#### try different bandwidth values
out1 = do.spufs(X, bandwidth=0.1)
out2 = do.spufs(X, bandwidth=1)
out3 = do.spufs(X, bandwidth=10)
#### visualize
opar <- par(no.readonly=TRUE)
par(mfrow=c(1,3))
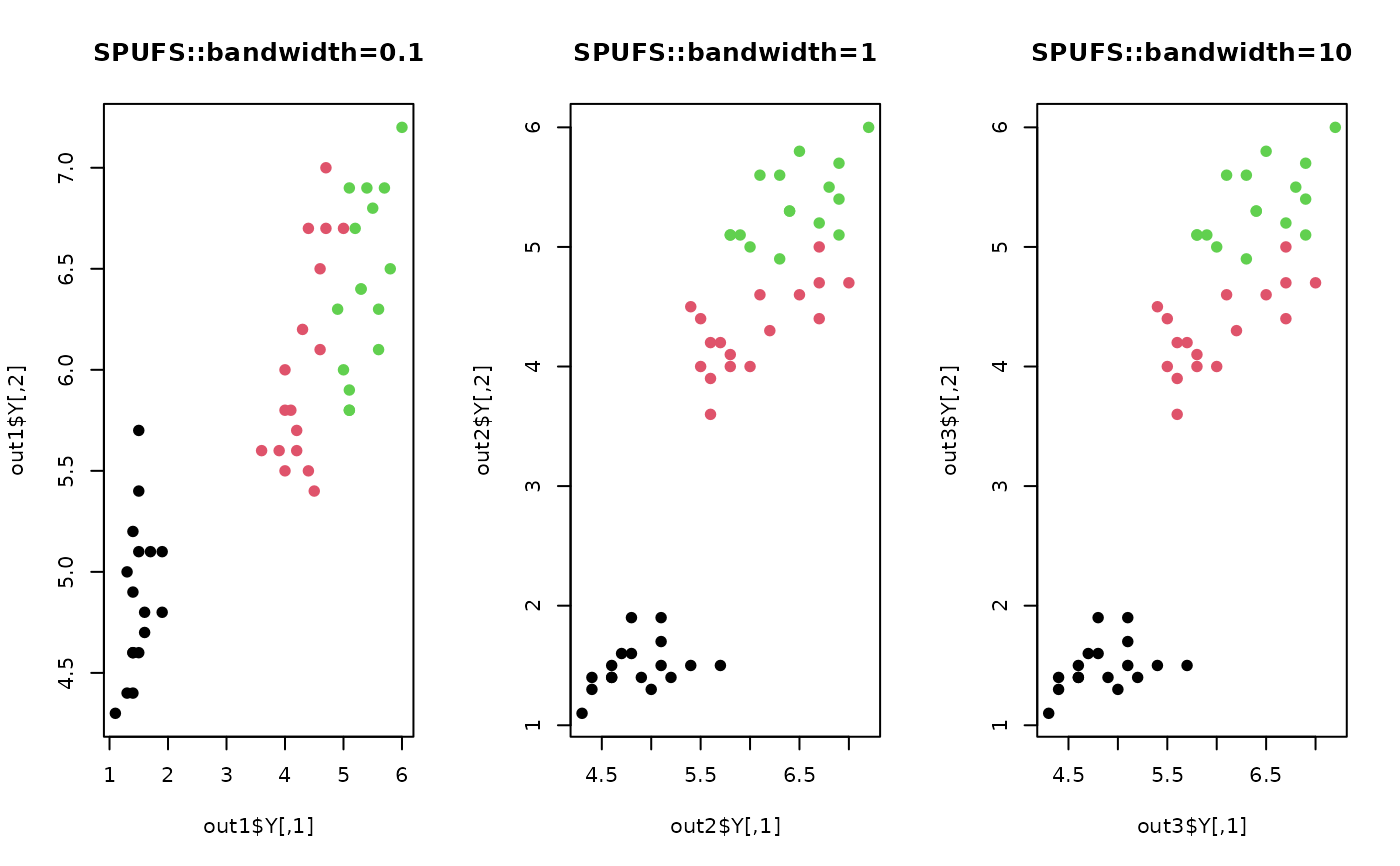
plot(out1$Y, pch=19, col=label, main="SPUFS::bandwidth=0.1")
plot(out2$Y, pch=19, col=label, main="SPUFS::bandwidth=1")
plot(out3$Y, pch=19, col=label, main="SPUFS::bandwidth=10")
 par(opar)
par(opar)